Input data and parameters
QualiMap command line
| qualimap bamqc -bam Map_with_BWA_MEM_on_dispersal_rep_3__and_data_26__mapped_reads_in_BAM_format_ -c -nw 400 -hm 3 -sd |
Alignment
| Command line: | bwa mem -t 5 -v 1 localref.fa /mnt/galaxy/files/001/447/dataset_1447881.dat /mnt/galaxy/files/001/448/dataset_1448087.dat |
| Draw chromosome limits: | yes |
| Analyze overlapping paired-end reads: | yes |
| Program: | bwa (0.7.17-r1188) |
| Analysis date: | Wed Jan 20 12:07:29 UTC 2021 |
| Size of a homopolymer: | 3 |
| Skip duplicate alignments: | yes (only flagged) |
| Number of windows: | 400 |
| BAM file: | Map_with_BWA_MEM_on_dispersal_rep_3__and_data_26__mapped_reads_in_BAM_format_ |
Summary
Warnings
| No flagged duplicates are detected | Make sure duplicate alignments are flagged in the BAM file or apply a different skip duplicates mode. |
Globals
| Reference size | 4,970,618 |
| Number of reads | 28,550,978 |
| Mapped reads | 27,786,327 / 97.32% |
| Unmapped reads | 764,651 / 2.68% |
| Mapped paired reads | 27,786,327 / 97.32% |
| Mapped reads, first in pair | 13,929,320 / 48.79% |
| Mapped reads, second in pair | 13,857,007 / 48.53% |
| Mapped reads, both in pair | 27,701,526 / 97.02% |
| Mapped reads, singletons | 84,801 / 0.3% |
| Secondary alignments | 0 |
| Supplementary alignments | 13,660 / 0.05% |
| Read min/max/mean length | 30 / 126 / 125.38 |
| Overlapping read pairs | 6,139,902 / 43.01% |
| Duplicated reads (flagged) | 0 / 0% |
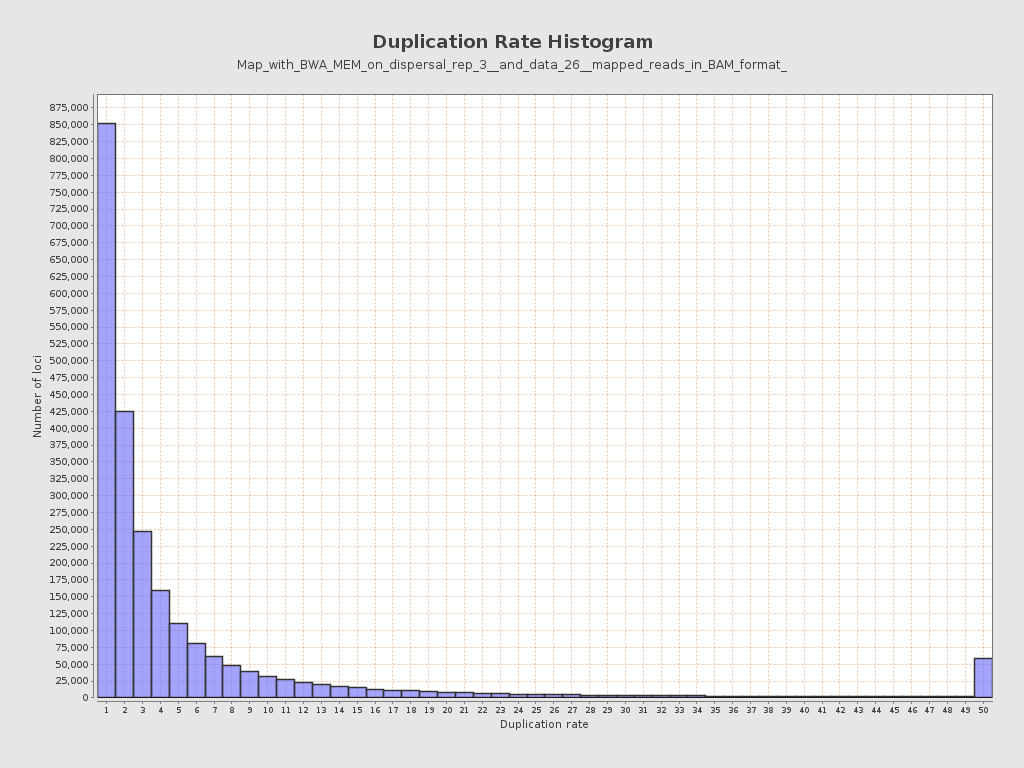
| Duplicated reads (estimated) | 25,444,873 / 89.12% |
| Duplication rate | 63.84% |
| Clipped reads | 13,266,425 / 46.47% |
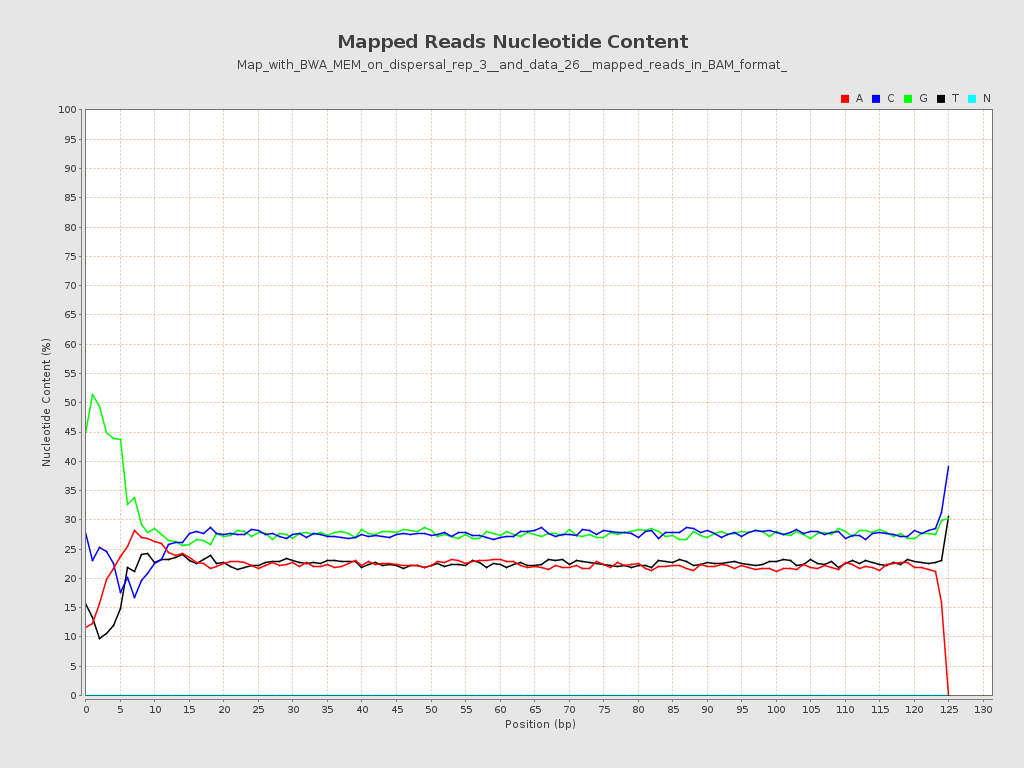
ACGT Content
| Number/percentage of A's | 745,869,297 / 22.16% |
| Number/percentage of C's | 914,152,964 / 27.16% |
| Number/percentage of T's | 748,205,227 / 22.23% |
| Number/percentage of G's | 957,421,863 / 28.45% |
| Number/percentage of N's | 201,056 / 0.01% |
| GC Percentage | 55.6% |
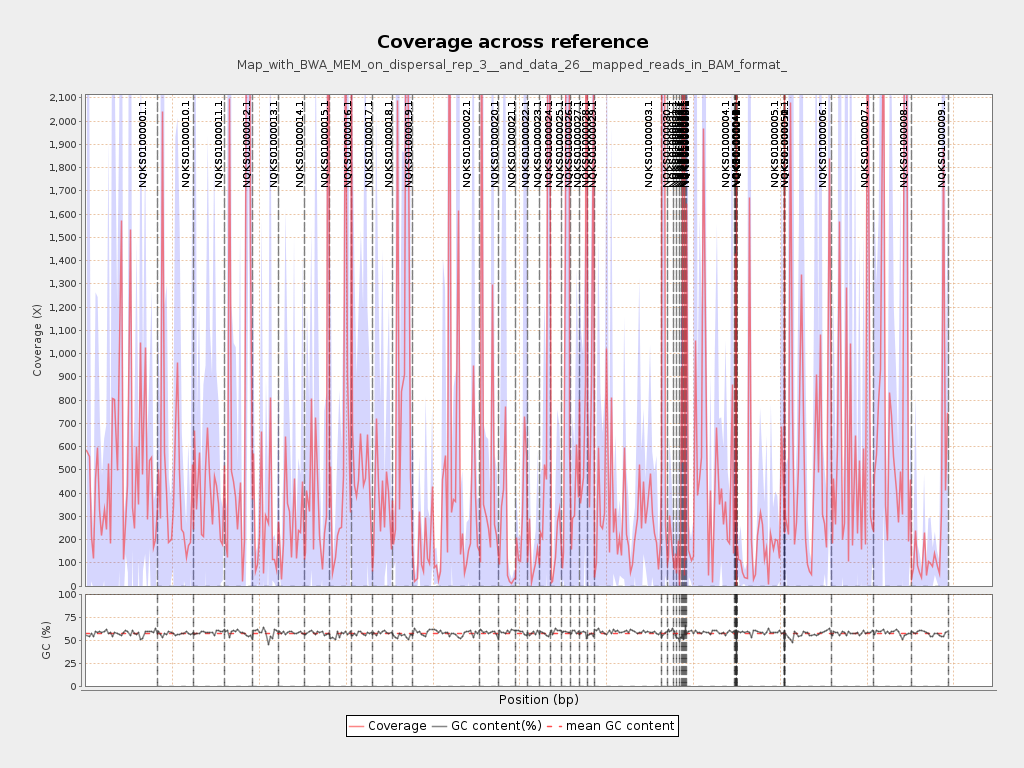
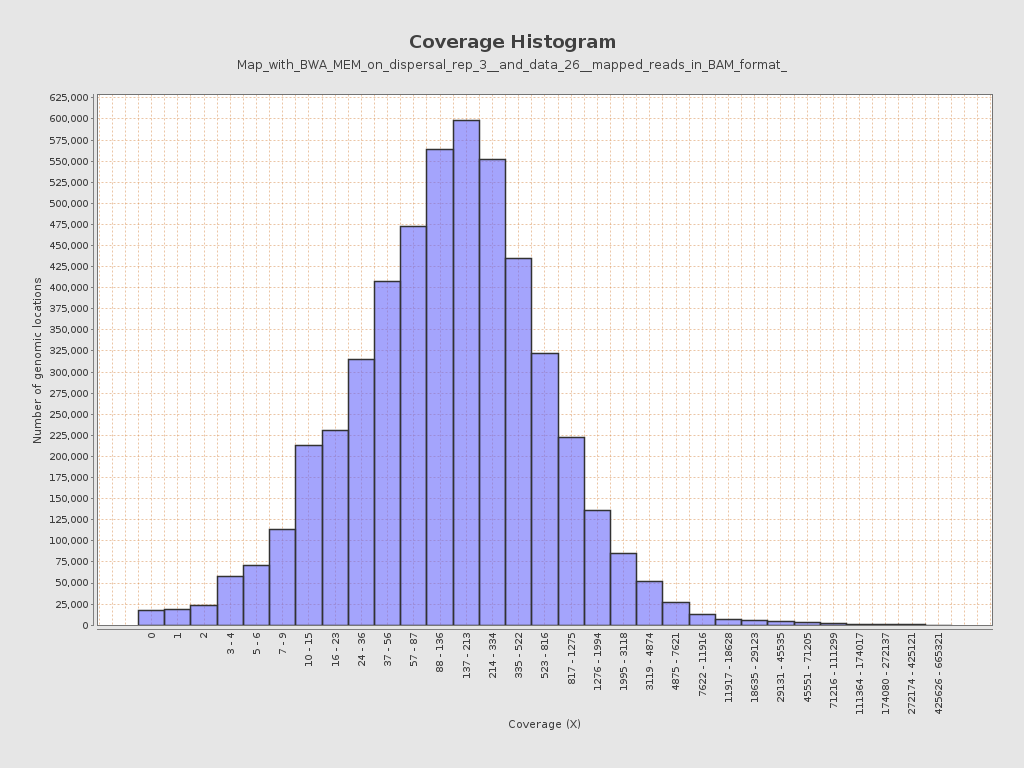
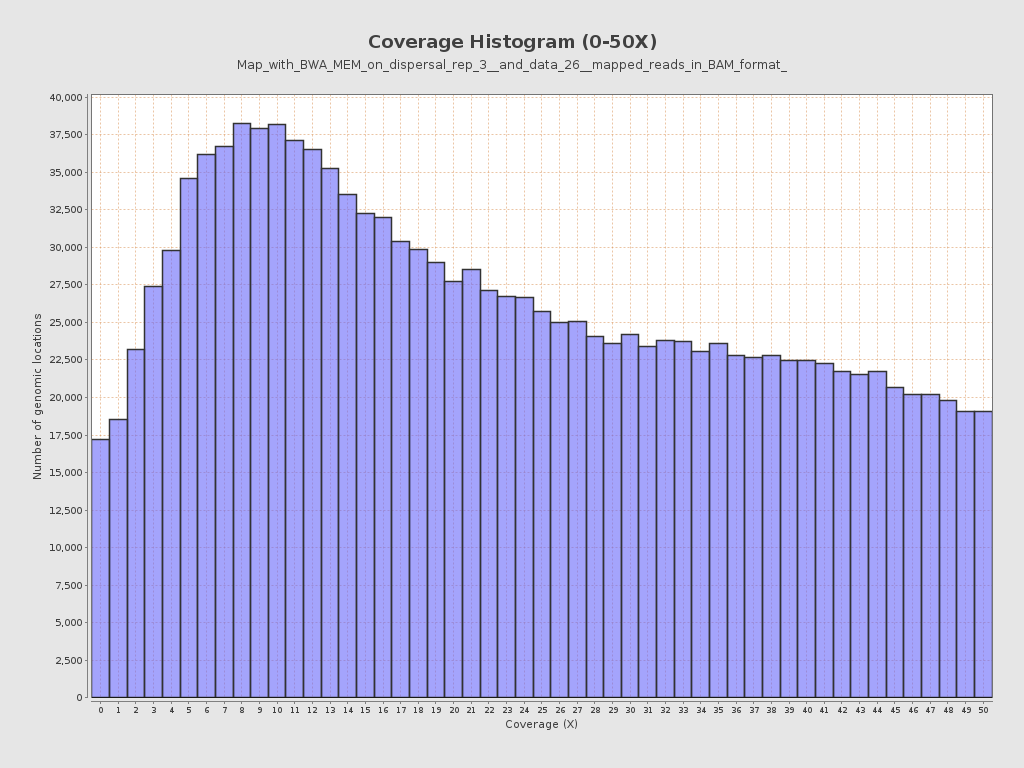
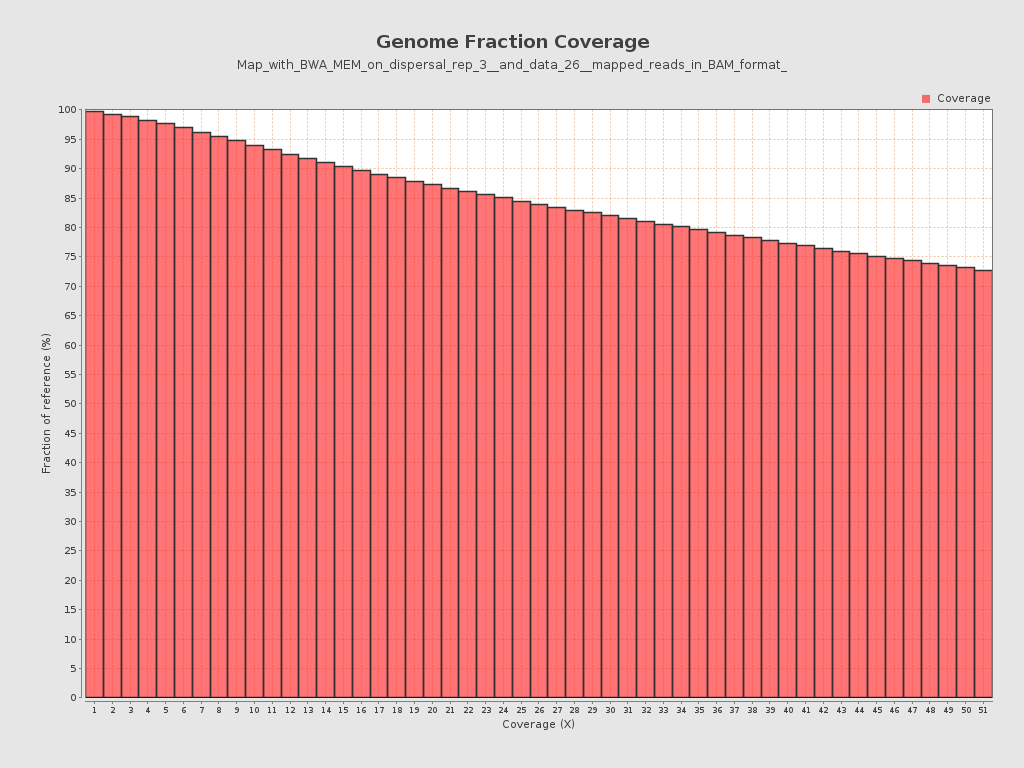
Coverage
| Mean | 677.1819 |
| Standard Deviation | 6,891.3243 |
| Mean (paired-end reads overlap ignored) | 618.99 |
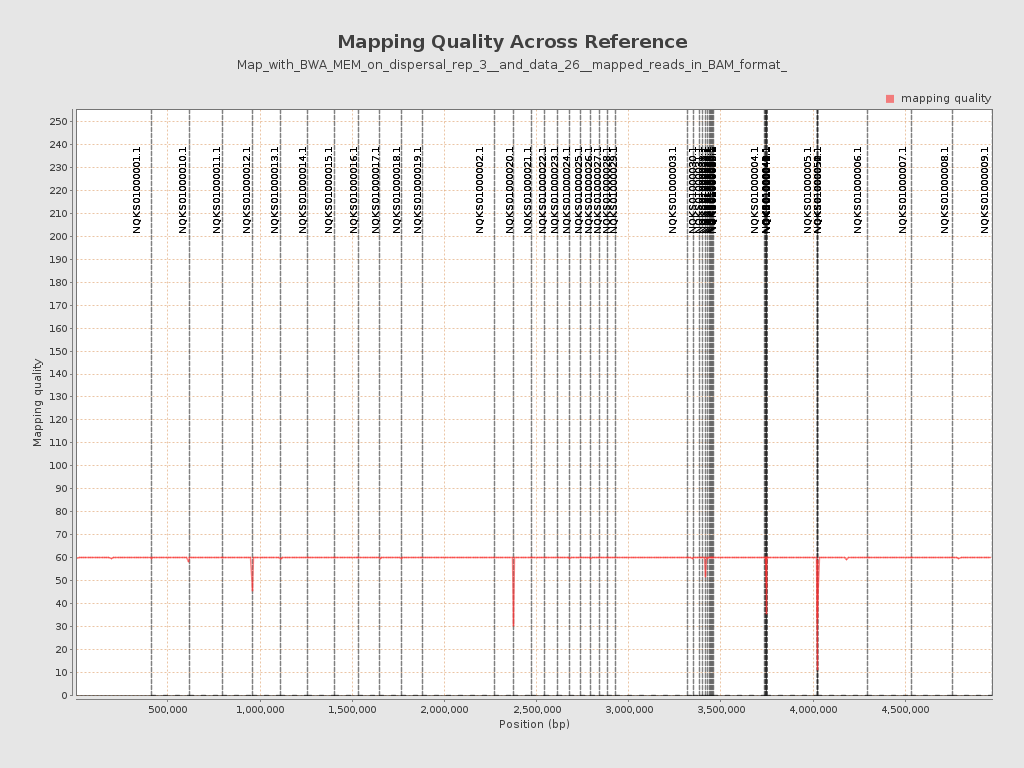
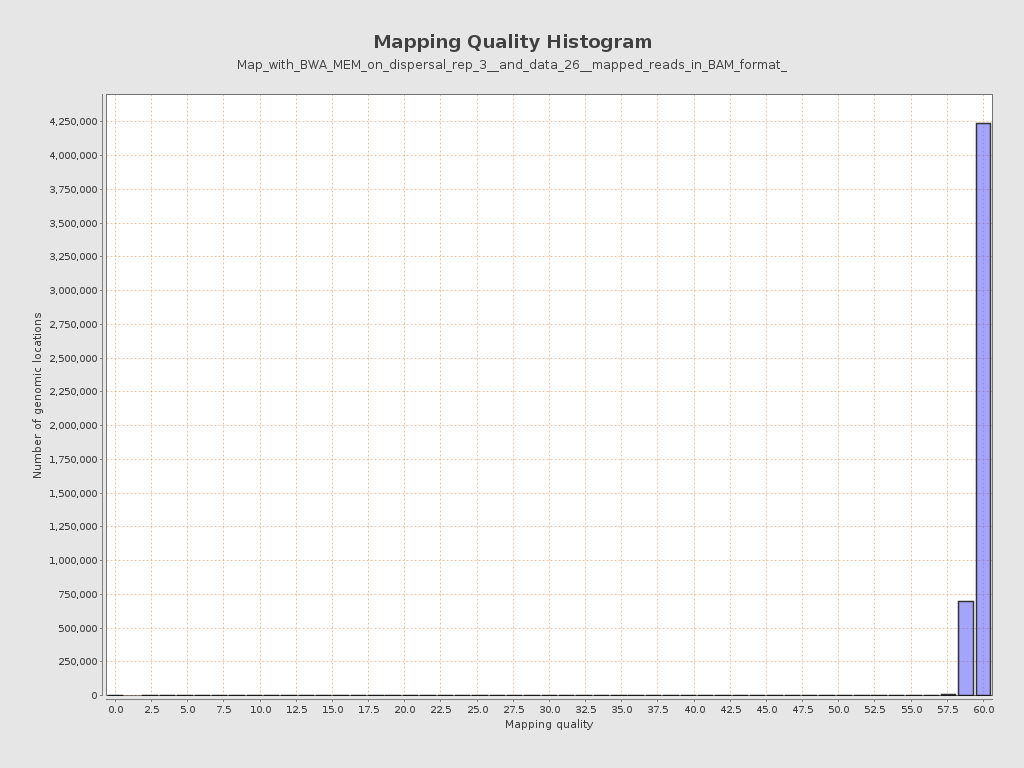
Mapping Quality
| Mean Mapping Quality | 59.56 |
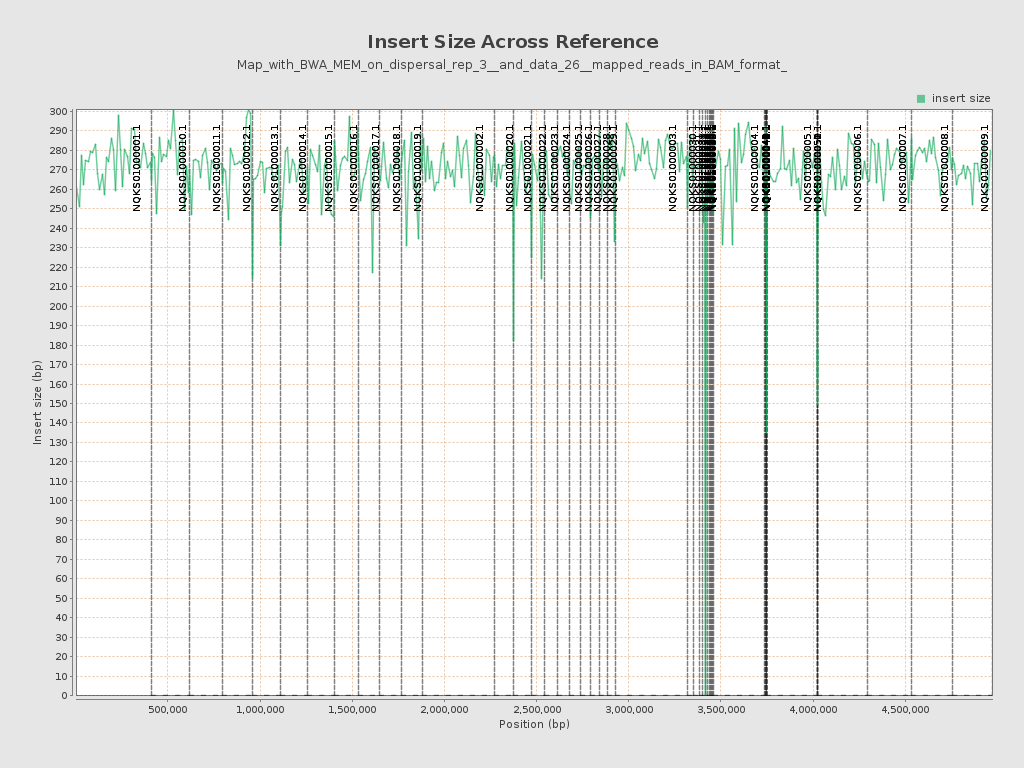
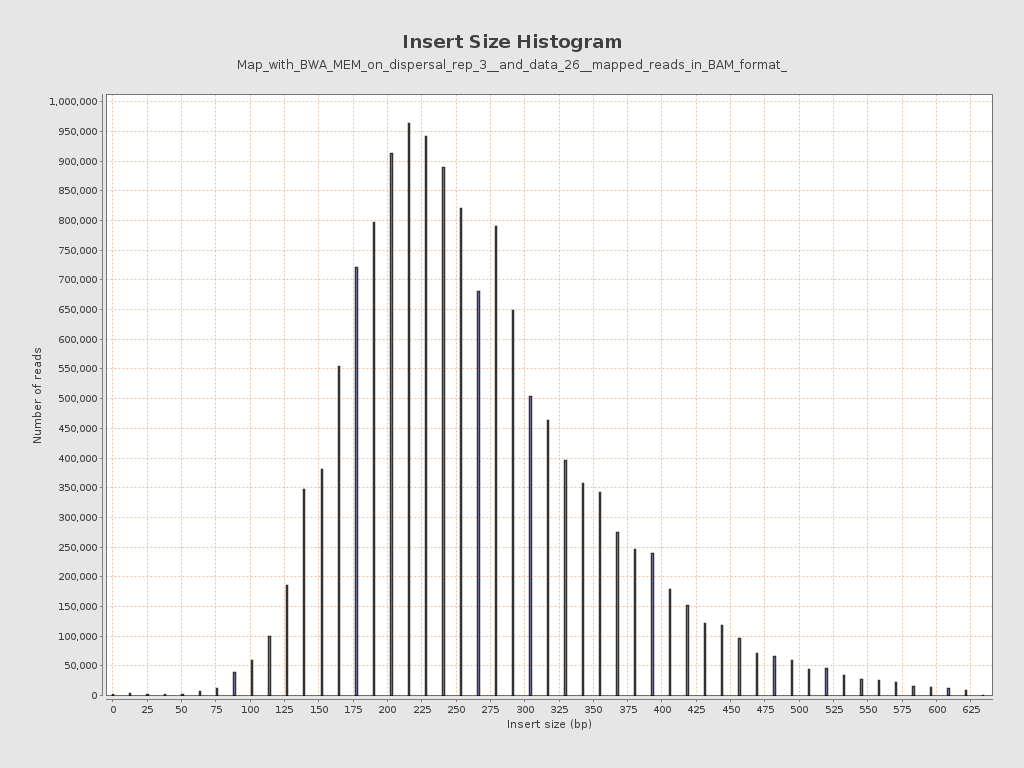
Insert size
| Mean | 271.18 |
| Standard Deviation | 228.57 |
| P25/Median/P75 | 206 / 253 / 317 |
Mismatches and indels
| General error rate | 0.32% |
| Mismatches | 10,561,228 |
| Insertions | 58,737 |
| Mapped reads with at least one insertion | 0.21% |
| Deletions | 63,149 |
| Mapped reads with at least one deletion | 0.21% |
| Homopolymer indels | 34.63% |
Chromosome stats
| Name | Length | Mapped bases | Mean coverage | Standard deviation |
| NQKS01000001.1 | 411870 | 215227309 | 522.5613 | 1,221.8496 |
| NQKS01000010.1 | 206580 | 99095696 | 479.6965 | 2,045.8686 |
| NQKS01000011.1 | 180011 | 71760103 | 398.6429 | 1,011.2717 |
| NQKS01000012.1 | 159844 | 161055645 | 1,007.5802 | 3,561.2431 |
| NQKS01000013.1 | 152608 | 42073441 | 275.6962 | 703.0423 |
| NQKS01000014.1 | 148176 | 41504102 | 280.1 | 596.5815 |
| NQKS01000015.1 | 142715 | 83840059 | 587.4649 | 4,067.7897 |
| NQKS01000016.1 | 130942 | 79175149 | 604.6582 | 3,270.1356 |
| NQKS01000017.1 | 117606 | 56028617 | 476.4095 | 1,002.1944 |
| NQKS01000018.1 | 115459 | 41533387 | 359.7241 | 745.2784 |
| NQKS01000019.1 | 113811 | 126203078 | 1,108.883 | 6,295.6581 |
| NQKS01000002.1 | 390567 | 136060358 | 348.3662 | 1,998.2684 |
| NQKS01000020.1 | 104542 | 69094277 | 660.9236 | 2,531.8609 |
| NQKS01000021.1 | 98658 | 21532495 | 218.2539 | 904.8389 |
| NQKS01000022.1 | 71460 | 23505836 | 328.937 | 1,553.0899 |
| NQKS01000023.1 | 69346 | 8430648 | 121.5737 | 282.6224 |
| NQKS01000024.1 | 63845 | 67867408 | 1,063.0027 | 6,931.9323 |
| NQKS01000025.1 | 61567 | 10715586 | 174.0476 | 322.0123 |
| NQKS01000026.1 | 53269 | 155569501 | 2,920.4509 | 9,362.1432 |
| NQKS01000027.1 | 48328 | 18480588 | 382.3992 | 538.6858 |
| NQKS01000028.1 | 45745 | 46543551 | 1,017.4566 | 4,568.8495 |
| NQKS01000029.1 | 41423 | 38670333 | 933.5474 | 2,444.4698 |
| NQKS01000003.1 | 387801 | 117585129 | 303.21 | 548.6701 |
| NQKS01000030.1 | 35809 | 107784643 | 3,009.9875 | 16,464.223 |
| NQKS01000031.1 | 31500 | 7066022 | 224.3182 | 279.0064 |
| NQKS01000032.1 | 18020 | 1914969 | 106.2691 | 146.5907 |
| NQKS01000033.1 | 15940 | 2653311 | 166.4561 | 383.3026 |
| NQKS01000034.1 | 11407 | 1303306 | 114.2549 | 255.1253 |
| NQKS01000035.1 | 9963 | 5729127 | 575.0403 | 574.9759 |
| NQKS01000036.1 | 5335 | 731227926 | 137,062.4041 | 132,935.5404 |
| NQKS01000037.1 | 5082 | 235122 | 46.2656 | 58.1791 |
| NQKS01000038.1 | 4561 | 5221435 | 1,144.8005 | 1,459.9811 |
| NQKS01000039.1 | 3508 | 1286039 | 366.6018 | 378.6035 |
| NQKS01000004.1 | 280988 | 137591013 | 489.6686 | 2,645.6074 |
| NQKS01000040.1 | 2821 | 1146933 | 406.5697 | 281.0486 |
| NQKS01000041.1 | 2595 | 1262476 | 486.5033 | 691.5879 |
| NQKS01000042.1 | 1560 | 2232250 | 1,430.9295 | 556.903 |
| NQKS01000043.1 | 1259 | 91654 | 72.799 | 57.4976 |
| NQKS01000044.1 | 373 | 197957 | 530.7158 | 290.1207 |
| NQKS01000045.1 | 371 | 11374790 | 30,659.8113 | 30,378.6433 |
| NQKS01000046.1 | 358 | 140205 | 391.6341 | 239.8282 |
| NQKS01000047.1 | 330 | 93955 | 284.7121 | 216.9591 |
| NQKS01000048.1 | 326 | 1328 | 4.0736 | 1.5231 |
| NQKS01000049.1 | 305 | 51447 | 168.6787 | 61.8032 |
| NQKS01000005.1 | 275949 | 66841789 | 242.2252 | 1,015.4667 |
| NQKS01000050.1 | 284 | 63916 | 225.0563 | 210.8725 |
| NQKS01000051.1 | 273 | 697883 | 2,556.348 | 2,337.9943 |
| NQKS01000052.1 | 244 | 85541 | 350.5779 | 262.7285 |
| NQKS01000006.1 | 271909 | 159935240 | 588.194 | 2,779.1316 |
| NQKS01000007.1 | 236695 | 143345386 | 605.6122 | 2,569.5826 |
| NQKS01000008.1 | 222895 | 186612656 | 837.2223 | 2,226.4549 |
| NQKS01000009.1 | 213785 | 58271792 | 272.5719 | 1,312.4071 |